Table of Contents
Abstract
Nonclinical contract research organizations (CROs) face a structural inefficiency in the preparation of exploratory and pivotal toxicology study packages: the Study Report and its companion SEND (Standard for Exchange of Nonclinical Data) dataset are generated through two separate, sequential processes from the same underlying data. This sequential workflow adds 4-5 weeks to submission timelines, duplicates significant data integration effort, and increases the risk of inconsistencies between the two deliverables. This paper describes a Single-Track process that merges these workflows, enabling concurrent preparation of the Study Report and SEND dataset from a unified data model. The approach reduces total package delivery time, eliminates redundant effort, and produces compliant-to-CDISC, consistent, higher quality submission-ready deliverables. The technology, process, and automation components required for successful implementation are described. Implications for CRO competitiveness, sponsor timelines, and regulatory review quality are discussed.
Background
Nonclinical CROs conduct animal safety studies on behalf of pharmaceutical, biotechnology, environmental and agrochemical sponsor clients. These studies span the full development continuum, from early exploratory dose-ranging and safety screening through complex, long-duration pivotal toxicology studies. Pivotal studies underpin Investigational New Drug (IND) filings, New Drug Applications (NDAs), and marketing authorizations worldwide.
At any given time, a single CRO manages a substantial number of studies simultaneously, spanning multiple study designs, species, and endpoints across regulatory jurisdictions. Clients range from small biotechs to large global pharma organizations, each with distinct needs and expectations.
Pivotal studies intended for regulatory submission are conducted under Good Laboratory Practice (GLP) standards, FDA (HHS, 21 CFR Part 58, PART 58—GOOD LABORATORY PRACTICE FOR NONCLINICAL LABORATORY STUDIES, 2026) in the United States, OECD GLP in Europe, and equivalent regional standards elsewhere. The primary deliverable is the Study Report, which documents the conduct, data, findings and conclusions of the study. Where mandated, a digitally standardized companion dataset – SEND (CDISC, 2021) – must also be submitted. Both deliverables are subject to rigorous quality standards, documentation controls, and audit trail requirements that assure regulatory agencies of data integrity, completeness, and faithful representation in the final Study Report. Meeting these standards demands intensive coordination across the Study Director’s team, data management, and Quality Assurance. The separate, sequential processes to get to the SEND and Study Report have been discussed (Rosentreter, 2018) as a cause for inconsistency.
Time, Quality, and Cost of the Current Sequential-Track Process
Both the Study Report and the SEND dataset must be derived from the same source data and must ultimately be consistent with one another. In practice, however, these deliverables are generated through two largely independent, sequential processes, a separation that has become an industry-wide norm since the SEND mandate took effect. The main process follows the study planning, followed by the study conduct and then the data analysis and study report drafting. The FDA as the early adopter of SDTM SEND standards, established the Technical Conformance Guide (HHS, 2025) that the CROs and their sponsor clients have to comply.
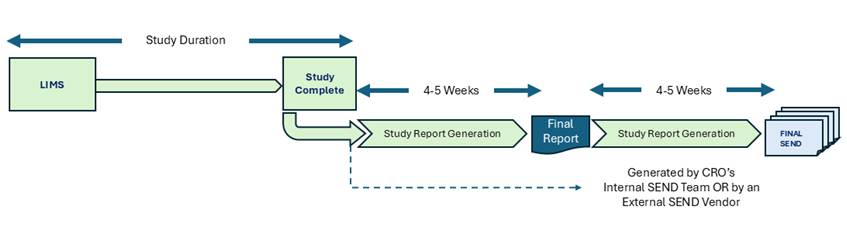
Figure: Current Sequential-Track Process for Nonclinical Studies at CROs
There is very little published about the current process used by Nonclinical Labs and CROs beyond the CFR Part 58 guidelines, and the individual CRO’s published SOPs. The perspective in (Thomas Steger-Hartmann et al, HHS, NIH, 2023) explains some of the challenges of integrating disparate data types of such studies. The current workflow operates sequentially, much like a relay race. The Study Director’s process is completed first until the Study Report is published. The SEND team does not begin substantive work until the Study Report is finalized by the Study Director. Their process does not have to follow GLP, but it must use the same GLP source as the one used by the Study Director. The TCG expects that the SEND data shall be capable of generating the same summary results as published in the Study Report. The cumulative consequence is a submission package that takes 8–10 weeks to complete: approximately 4–5 weeks for the Study Report after data completion, followed by another 4–5 weeks for SEND generation and quality assurance checks. This structure imposes compounding costs across three dimensions: timeline, data integration effort, and quality assurance.
1. Timeline Impact
- SEND generation begins only after the Study Report is complete, adding its full cycle time sequentially behind the Study Report generating process, rather than concurrently.
- Total submission package time: 8–10 weeks, versus the 4–5 weeks theoretically required if the two processes were done in a single, merged process.
2. Duplicated Data Integration Effort
- Protocol metadata extraction is performed twice. Key study design parameters, treatment groups, dosing regimens, species, and route of administration, must be extracted from the negotiated Protocol (along with the CRO’s Study Plan), and its amendments, first for the Study Director’s use, then again by the SEND team to populate the Trial Summary (TS), Trial Design (TE, TA, TX), and Exposure (EX) domains.
- Non-LIMS data integration is duplicated. The Study Director’s team must compile data from the LIMS alongside non-LIMS sources, including bioanalysis, ADA, FACS, immunology markers, multi-omics, and other ancillary findings. The SEND team must then independently re-integrate this same data. Automated LIMS adaptors, sometimes cited as a mitigating factor, do not address non-LIMS data sources and therefore do not reduce this integration cost.
- Study Report metadata must be re-extracted for SEND. A high proportion of Trial Summary parameters (TS.XPT) are captured in the Study Report methods sections and facility sub-reports, not in the LIMS. These must be manually re-extracted by the SEND team, representing a further duplication of effort.
3. Quality Assurance Costs
- Disjointed processes create unnecessary errors. Running two disjointed processes from the same source data by separate teams creates structural conditions for divergence. The Study Report and SEND dataset have more opportunities to become inconsistent with one another than to remain aligned by chance.
- Rigorous reconciliation is resource-intensive. Mechanically reconciling the SEND dataset against the Study Report, including digitizing the report, extracting tabulations, and harmonizing terminology to CDISC controlled terminology, accounts for approximately 30% or more of total SEND generation cost. Digitization of the Study Report alone represents roughly 20% of this figure. Where Study Report appendices include data domains not captured in LIMS or non-LIMS sources, reconciliation costs can approach 40%.
- Spot-checking is the prevailing substitute, and is insufficient. CROs and SEND vendors rely on manual spot-checks of tabulated SEND data against Study Report PDF appendices rather than systematic reconciliation. This approach is both costly and non-rigorous, leaving substantive inconsistencies undetected.
- Upstream disorganization compounds downstream cost. A significant portion of reconciliation effort, approximately 80% of the quality cost component, arises from reconstituting subject-level SEND data into group summaries and harmonizing extracted tabulation terminology to CDISC controlled terminology prior to comparison.
The Single-Track Process: Merged Study Report and SEND Process
The Single-Track approach merges Study Report authoring and SEND dataset preparation into a unified workflow, operating from a shared, continuously updated data model. Rather than treating the Study Report as a prerequisite for SEND, both deliverables are developed in parallel from the same integrated data layer, with the SEND dataset finalized at the moment the Study Report is marked Final. One dataset, two deliverables.
Critically, the Single-Track design does not impose new cross-functional dependencies. Study Directors are not required to learn SDTM, CDISC, or SEND controlled terminology, and the SEND team’s workflow is not made dependent on the Study Director’s schedule. Both organizations continue to work within their established practices; the integration occurs at the data and metadata layer, not at the organizational level.
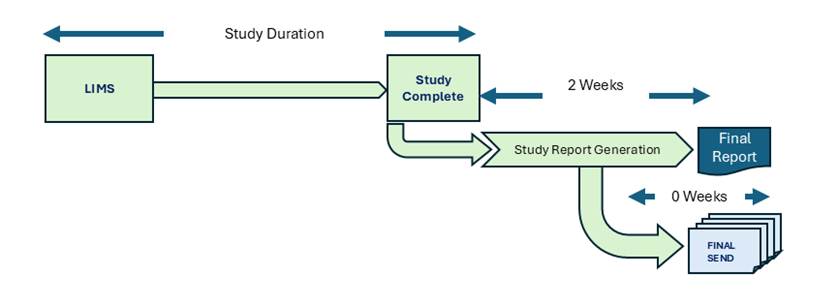
Figure: Single-Track Process for Nonclinical Studies at CROs
1. Study Report Authoring
As study data are collected and ingested, the Study Director’s team curates them within a Universal Data Model (UDM), producing an analysis-ready, integrated view of the study data through a Data Services Layer (DSL). Dose- and time-related signals are computed using fold-change analysis and automated statistical tests, generating a concise set of potentially noteworthy findings. Concordance analyses spanning clinical pathology, histopathology, organ macroscopy, in-vivo measurements, and clinical observations further distill these signals, providing the Study Director with a structured basis for final assessments and sponsor reviews. This integrated view encompasses LIMS data alongside all non-LIMS sources, bioanalysis, ADA, FACS, immunology, and ancillary findings, within a single coherent structure.
Driven by the target Study Report Table of Contents, maintained within a metadata registry, the Study Director is provided with AI-assisted, section-by-section drafting of report narrative. Draft text is generated from the integrated study data, computed dose-response and time-course signals, protocol content, and relevant facility logs. The Study Director may prompt the AI to refine and shape the text or edit draft sections directly. The system automatically validates edited text against the underlying data to ensure that no material change in meaning occurs relative to the data-supported findings.
Once final report draft reviews are complete, the study data are locked and versioned in conjunction with the Study Report. All subsequent modifications are subject to 21 CFR Part 11 electronic records controls.
2. Contemporaneous SEND Metadata Capture
In parallel with Study Report authoring, the SEND team captures metadata prospectively, beginning before Study Start. Key protocol parameters are entered directly into the SEND Trial Summary domain editor, enabling generation of the TE, TA, TX, and EX domains from Protocol content prior to first in-life procedures. This prospective capture eliminates the need for retroactive extraction and re-entry.
As the study progresses, LIMS data are read continuously by the integrated process to populate the DM (Demographics) domain. The Study Director’s team’s decisions regarding methods, instruments, and practices are captured in the Metadata Registry, the authoritative internal reference for standards, methods, and instruments, and automatically reflected in the SEND metadata layer. This eliminates the separate manual extraction step that characterizes the current sequential-track process.
3. SEND Quality Control and Submission Packaging
Once study data are fully ingested, the SEND QC team may preview the dataset at any time, generating test exports to the selected Implementation Guide (IG) and controlled terminology (CT) version. Validation reports and annotations accumulated during metadata entry and review are automatically compiled into the nonclinical Study Data Reviewer’s Guide (nSDRG). The Define.XML is automatically generated by the transformer, drawing on the CDISC standards and controlled terminologies held in the metadata registries.
When the Study Report is marked Final, the SEND dataset is immediately exportable as a submission-ready package, including SEND datasets, nSDRG, Define.XML, and validation reports, without additional cycle time.
Essential Technology, Process, and Automation Components
The Single-Track approach is transformative for CROs. Faster study deliveries, cost reductions of 5%-10% per study, and quality and consistency assured by the single-track design across both deliverables transforms the CRO’s business to make it more agile and competitive. Realizing these benefits, however, requires the deployment of three interdependent capability sets:
- Technology,
- Process, and
- Automation.
These cannot be just improvised or assembled informally. Each must be proven across the full range of study data types and maturation stages, from in vivo data collection through Study Report finalization and SEND submission. The absence or inadequate implementation of any one of these three capabilities will increase the labor, cost and time for implementing a Single-Track workflow.
1. Technology
Six core technology components underpin the Single-Track process. Together, they constitute an integrated platform rather than a collection of point solutions.
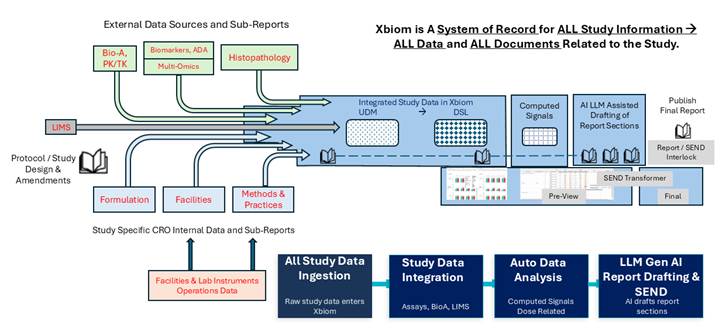
Figure: Single-Track Enabled Xbiom™ Study Information Management System
-
Exchange Portal. CROs need to collaborate with sponsor clients across the full lifecycle of a study, from protocol receipt through final draft reviews and deliverable acceptance. They also need to collaborate with their network of specialist vendors and contract laboratories that contribute assay data for their studies. The Exchange Portal provides a purpose-built, out-of-the-box collaboration environment with end-to-end encryption (data in transit and at rest), automated crawlers, and event-driven workflow triggers. It replaces ad-hoc email and file transfer with governed, auditable data exchange.
- Layered Data Repository. The data repository is the most architecturally critical component of the Single-Track platform. It departs from the file-based data management traditions that prevail in the industry and instead employs large, elastic data stores capable of indexing and co-locating data of all types, from structured relational records to streaming instrument inputs to unstructured document content, at scale, with regulatory compliance built in.
The repository is organized into five progressive layers, with a sixth that is relevant to BigPharma who are building large tensors for predictive safety using neural network for deep learning. These layers are populated by automated maturation processes moving data from each layer to the next:
-
As-Collected Data. Raw study data from all sources, including structured LIMS outputs, streaming instrument data, and unstructured content such as pathology narratives and facility logs.
- Metadata Layer. Study-level and standards metadata, including protocol parameters, CDISC controlled terminology mappings, and facility-specific reference standards held in the Metadata Registry.
- Data Services Layer (DSL). Analysis-ready data, curated and harmonized from all sources. This layer serves as the authoritative integrated view of the study for the Study Director’s team.
- Derived Signals and Endpoints. Computed dose- and time-related signals, statistical test outputs, tabulations, figures, and listings (TFLs) supporting report sections for summaries, results and conclusions.
- Concordance and Reportable Signal Vector. Cross-domain concordance outputs and distilled signal vectors presented to the Principal Investigator and Study Director for narrative review and assessment of final set of noteworthy reportable safety signals.
- Document and Content Management System (DMS). An integrated DMS that operates in concert with the Data Repository and Metadata Registry, managing the full lifecycle of study documents, from protocol receipt and amendment tracking through draft report review, approval workflows, and final archival. The DMS provides the governance layer for all human-in-the-loop content review and sign-off.
- Metadata Registry (MDR). The MDR is the governance backbone of the Single-Track platform. It maintains the authoritative definitions of all standards, controlled terminologies, regulatory rules, and corporate guidelines that govern study conduct, reporting, and SEND generation. The MDR ensures that all processes, automated and human, operate within defined compliance guardrails. Attempting to implement Single-Track without a robust MDR, for example using spreadsheet-based approaches, is not viable: the complexity and interdependency of the metadata relationships required cannot be managed in unstructured formats.
- Transformer. The Transformer is a machine-learning-enabled, low-code ETL (extract, transform, load) engine that converts data between any source and any target model using mapping rules held in the MDR. It performs the bidirectional transformations required to decompose PDF documents into their constituent components and to assemble structured content into the Study Report Table of Contents structure. The Transformer is the engine of automated SEND generation. Once the Study Director locks the study data, the Transformer produces submission-ready SEND XPT or JSON files directly from the versioned data layer, without manual intervention.
-
AI Large Language Model (LLM). Large Language Models capable of tokenizing document content, interpreting its meaning, and generating contextually accurate narrative text from structured data signals are a key component of the Study Report authoring workflow. Critically, LLMs are not integrated directly into the data or content management layers. Instead, they interact with the Single-Track platform exclusively through read-only APIs or Retrieval-Augmented Generation (RAG) interfaces, ensuring that the LLM cannot modify source data or metadata. All LLM-generated text passes through a human-in-the-loop review process governed by the Single-Track solution before being deposited in the DMS. Autonomous or agentic LLM workflows are deliberately avoided: LLMs are employed for data-to-text and text-to-data conversion tasks only, where their outputs can be validated against the underlying data with full traceability. The guiding principles (FDA, 2026) and guard rails around the application of AI LLM are discussed in a recent publication by FDA and EMA.
- Supporting Technology Components. A number of additional capabilities complete the platform: scripting environments (R, Python) for custom statistical computations; interactive data, metadata, and signal visualization with integrated annotation tools; automated validation against CDISC, FDA, and PMDA rules; OCR and LLM-augmented OCR for structured data extraction from PDF content; and email integration to automate and archive all study-related communications as part of the regulatory record.
2. Process
The Single-Track workflow is organized around a defined set of processes that govern how data mature from collection through final submission. Two categories of process are applicable.
- Automated Maturation Processes. These processes run autonomously or are triggered by data events, elapsed time, or predefined rules. They advance the study data through successive layers of the repository, from as-collected data through preparation, analysis, and signal generation, and are punctuated by defined toll gates or checkpoints that require human review or sign-off before the study advances. Examples include automated ingestion of LIMS data into the DSL, triggered generation of dose-response signals when endpoint data are received, and automated reading of facility logs with highlighting of dose-related observations for the Study Director.
- Tethered and Event-Driven Workflows. These workflows are triggered by upstream process completion or by manual initiation, and operate alongside, but not in the critical path of, the primary study maturation processes. Examples include data exchange crawlers, sponsor collaboration notifications, and draft report review routing. While not critical-path, these workflows reduce administrative overhead and ensure that all relevant parties are informed and engaged at appropriate points in the study lifecycle.
3. Automation
Automation is the mechanism by which the Single-Track approach achieves its efficiency gains. Two forms of automation are employed, with distinct roles and constraints.
- Rule-Based and Triggered Automation. Deterministic, triggered automation is used for activities where precision, traceability, and regulatory compliance are paramount. SEND dataset generation, signal computation, facility log ingestion, and Define.XML production are all executed through rule-based automation governed by the MDR. These processes produce fully auditable outputs with complete lineage from source data to final deliverable.
- AI-Assisted Automation and Agentic Boundaries. AI-assisted automation, including LLM-based data-to-text and text-to-data conversion, is used where judgment, natural language interpretation, or narrative generation is required. However, fully autonomous agentic AI workflows are deliberately excluded from the Single-Track implementation. Current LLM capabilities do not provide the level of deterministic rigor and traceability required for GLP-compliant study reporting. LLMs are therefore constrained to advisory and generative roles, with all outputs subject to human-in-the-loop review before finalization. This boundary preserves regulatory compliance while capturing the productivity benefits of AI assistance.
Summary of Benefits
The Single-Track process delivers measurable improvements across the three dimensions that matter most to CROs and their sponsor clients:
- Time: Total submission package cycle time is reduced from 8–10 weeks to approximately 4–5 weeks by eliminating the separate, sequential dependency between Study Report and SEND generation.
- Cost: Duplicated data integration, protocol metadata extraction, and reconciliation effort are eliminated or substantially reduced. Savings in SEND generation, which ranges from 5%-10% of the Study cost, with the 5%-8% savings in the Study Director’s workflow leading to the Study Report.
- Quality: Structural consistency between the Study Report and SEND dataset is assured by their derivation from a single, locked, versioned study data model. Systematic reconciliation or spot-checking with their discrepancies are now resolved at source rather than detected post-finalization of the two deliverables.
Conclusion
The sequential-track model persists largely out of convention, not because it serves CROs or their clients well. Adding 4–5 weeks of sequential SEND work after an already intensive reporting cycle is a cost that compounds across a portfolio of studies, and the reconciliation effort required to verify consistency between two independently produced deliverables absorbs resources without adding scientific value. The Single-Track process eliminates this sequential dependency. By deriving both the Study Report and SEND dataset from a single versioned data model, the submission package is complete when the Study Director signs off, not weeks later. The practical result is that CROs can turn around studies faster without adding headcount, sponsors see shorter paths to IND and NDA filing, and reviewers at FDA or PMDA receive packages where the narrative and the underlying dataset actually agree with one another by construction rather than by post-hoc checking.
Singe-Track requires a data platform meant for studies, with disciplined metadata governance, and careful boundaries around where automation can and cannot replace human oversight. The components described in this paper are proven individually, the challenge is now integration and operational commitment, not invention.
Key Abbreviations
| ADA | Anti-Drug Antibody |
| AI | Artificial Intelligence |
| CDISC | Clinical Data Interchange Standards Consortium |
| CFR | Code of Federal Regulations |
| CRO | Contract Research Organization |
| CT | Controlled Terminology |
| DMS | Document and Content Management System |
| DSL | Data Services Layer |
| ETL | Extract, Transform, Load |
| EX | Exposure (SEND domain) |
| FACS | Fluorescence-Activated Cell Sorting |
| FDA | U.S. Food and Drug Administration |
| GLP | Good Laboratory Practice |
| IG | Implementation Guide |
| IND | Investigational New Drug |
| LIMS | Laboratory Information Management System |
| LLM | Large Language Model |
| MDR | Metadata Registry |
| NDA | New Drug Application |
| nSDRG | Nonclinical Study Data Reviewer’s Guide |
| OCR | Optical Character Recognition |
| OECD | Organization for Economic Co-operation and Development |
| PMDA | Pharmaceuticals and Medical Devices Agency (Japan) |
| QC | Quality Control |
| RAG | Retrieval-Augmented Generation |
| SDTM | Study Data Tabulation Model |
| SEND | Standard for Exchange of Nonclinical Data |
| SOP | Standard Operating Procedure |
| TA | Trial Arms (SEND domain) |
| TCG | Technical Conformance Guide |
| TE | Trial Elements (SEND domain) |
| TFL | Table, Figure, and Listing |
| TS | Trial Summary (SEND domain) |
| TX | Trial Set (SEND domain) |
| UDM | Universal Data Model |
| XPT | SAS Transport File Format |
Bibliography
CDISC. (2021). CDISC SEND Implementation Guide. Retrieved from https://www.cdisc.org/standards/foundational/send
FDA, E. (2026, Jan). Guiding Principles of Good AI Practice in Drug Development. Retrieved from https://www.fda.gov/media/189581/download
HHS, F. (n.d.). Retrieved from https://www.fda.gov/media/153632/download
HHS, F. (2025). STUDY DATA TECHNICAL CONFORMANCE GUIDE. Retrieved from https://www.fda.gov/media/153632/download
HHS, F. (2026). 21 CFR Part 58, PART 58—GOOD LABORATORY PRACTICE FOR NONCLINICAL LABORATORY STUDIES. Retrieved from https://www.ecfr.gov/current/title-21/chapter-I/subchapter-A/part-58
Rosentreter, M. a. (2018). Data Consistency – SEND Datasets and Study Reports: Request for Collaboration in the Dynamic SEND Landscape. PhUSE.
Thomas Steger-Hartmann et al, HHS, NIH. (2023). Perspectives of data science in preclinical safety assessment. Retrieved from Drug Discovery Today: https://pubmed.ncbi.nlm.nih.gov/37244565/
[1] PointCross Life Sciences Inc., [email protected]
[2] ToxiStrategy, [email protected]
[3] PointCross Life Sciences Inc., [email protected]
[4] PointCross Life Sciences Inc., [email protected]

